Application of Fourier Methods to Computer Simulation of Supercoiled DNA
Fourier methods have been applied to model the
overall geometry of a supercoiled DNA curve, the starting coordinates of
which can be obtained from electron microscopy measurements or other theoretical
simulations. Evenly spaced points on a known DNA curve are selected and
subsequently used to model the original curve in terms of a finite Fourier
series. Hence, a simple analytical curve expression, which closely resembles
the initial data, is obtained for analysis and optimization.
Energy minimization of DNA supercoils represented by
such finite Fourier series has been performed using an elastic energy model.
Minimized configurations are identified for interwound supercoils of 1000
bp at various values of the linking number difference,  Lk.
The configurational profiles of writhing numbers (Wr) and energy
vs.
Lk.
The configurational profiles of writhing numbers (Wr) and energy
vs.  Lk are similar to those of
optimized structures previously found with a B-spline representation of
the DNA supercoil. However there are some notable differences that probably
result from the more global nature of the Fourier modeling compared
to the B-spline. At the same
Lk are similar to those of
optimized structures previously found with a B-spline representation of
the DNA supercoil. However there are some notable differences that probably
result from the more global nature of the Fourier modeling compared
to the B-spline. At the same  Lk,
a higher value of the writhing number and a lower energy are consistently
obtained with the finite Fourier series representation. Unlike the
B-spline minimized structures and the configurations identified earlier
by finite element analysis, only two families, the figure-eight and the
loose interwound forms at low
Lk,
a higher value of the writhing number and a lower energy are consistently
obtained with the finite Fourier series representation. Unlike the
B-spline minimized structures and the configurations identified earlier
by finite element analysis, only two families, the figure-eight and the
loose interwound forms at low  Lk,
are found to overlap; there is no overlapping of interwound configurational
families at higher
Lk,
are found to overlap; there is no overlapping of interwound configurational
families at higher  Lk.
Lk.
The optimized configurations of three-lobed 1000 bp branched DNA supercoils
have also been identified. Different families of structures are found over
a /\Lk range between -0.6 and 6.0. Family I, with three lobes of
similar shape and size and with Wr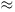 0,
occurs at low
0,
occurs at low  Lk values.
Families II-VI, also with three lobes of similar shape and size, exist
over a range of
Lk values.
Families II-VI, also with three lobes of similar shape and size, exist
over a range of  Lk from 1.0
to 4.6. The structures in families IIa-Va with one lobe larger than the
others have similar Wr values at the same
Lk from 1.0
to 4.6. The structures in families IIa-Va with one lobe larger than the
others have similar Wr values at the same  Lk
but are slightly higher in energy. Families VII and VIII of minima
found between
Lk
but are slightly higher in energy. Families VII and VIII of minima
found between  Lk of 3.2 and
6.0 are characterized by one interwound and two open lobes, the later being
similar to but smaller than the lobes in the simpler branched structures
of families II-VI. The occurrence of branched interwound energy minima
is consistent with the highly branched configurations of supercoiled DNA
observed under the electron microscope.
Lk of 3.2 and
6.0 are characterized by one interwound and two open lobes, the later being
similar to but smaller than the lobes in the simpler branched structures
of families II-VI. The occurrence of branched interwound energy minima
is consistent with the highly branched configurations of supercoiled DNA
observed under the electron microscope.
Configurational snapshots of long partially relaxed DNA plasmids (6400
bp) taken from electron microscopic images are found to optimize to closely
related linear and branched interwound forms with writhing numbers comparable
to those of the nearly planar projected starting states. The energy minimization
tends to straighten and maximize nonbonded contacts between interwound
double helical strands and to fix the size of the hairpin loops emerging
from the interwound domains. The configurational similarities of
these minimized "random" starting states with the families of optimized
configurations systematically identified at shorter chain lengths affirm
the completeness of the search for energy minima.

 Click to go back to the publication list
Click to go back to the publication list
Webpage design by Igor Zilberman

 Click to go back to the publication list
Click to go back to the publication list