Resonance Analysis in Force Splitting Methods for Biomolecular Dynamics
Numerical resonance artifacts have become recognized recently as a limiting
factor to increasing the timestep in multiple-timestep (MTS) biomolecular
dynamics simulations. At certain timesteps correlated to internal motions
(e.g., 5 fs, around half the period of the fastest bond stretch, Tmin),
visible inaccuracies or instabilities can occur. Impulse-MTS schemes are
vulnerable to these resonance errors since large energy pulses are introduced
to the governing dynamics equations when the slow forces are evaluated.
We recently showed that such resonance artifacts can be masked significantly
by applying extrapolative splitting to stochastic dynamics. Theoretical
and numerical analyses of force-splitting integrators based on the Verlet
discretization are reported here for linear models to explain these observations
and suggest how to construct effective integrators for bimolecular dynamics
that balance stability with accuracy.
Analyses for Newtonian dynamics demonstrate the severe resonance patterns
of the Impulse splitting, with this severity worsening with the outer timestep,  t;
Constant Extrapolation is generally unstable, but the disturbances
do not grow with
t;
Constant Extrapolation is generally unstable, but the disturbances
do not grow with  t. Thus,
the stochastic extrapolative combination can counteract generic instabilities
and largely alleviate resonances with a sufficiently strong Langevin heat-bath
coupling (
t. Thus,
the stochastic extrapolative combination can counteract generic instabilities
and largely alleviate resonances with a sufficiently strong Langevin heat-bath
coupling (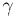 ), estimates for which
are derived here based on the fastest and the slowest motion periods. These
resonance results generally hold for nonlinear test systems: a water tetrameter
and solvated protein. Proposed related approaches such as Extrapolation/Correction
and Midpoint Extrapolation only work better than Constant Extrapolation
for timesteps less than Tmin/2.
), estimates for which
are derived here based on the fastest and the slowest motion periods. These
resonance results generally hold for nonlinear test systems: a water tetrameter
and solvated protein. Proposed related approaches such as Extrapolation/Correction
and Midpoint Extrapolation only work better than Constant Extrapolation
for timesteps less than Tmin/2.
An effective extrapolative stochastic approach for biomolecules that balances
long-timestep stability with good accuracy for the fast subsystem
is then applied to a biomolecule using a three-class partitioning: the
medium forces are treated by Midpoint Extrapolation via position
Verlet, and the slow forces are incorporated by Constant Extrapolation.
The resulting algorithm (LN) performs well on a solvated protein system
in terms of themodynamical properties and yields an order of magnitude
speedup with respect to single-timestep Langevin trajectories. Computed
spectral density functions also show how the Newtonian modes can be approximated
by using a small 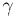 in the range
of 5-20 ps-1.
in the range
of 5-20 ps-1.

 Click to go back to the publication list
Click to go back to the publication list
Webpage design by Igor Zilberman

 Click to go back to the publication list
Click to go back to the publication list