Tree Topology: 4_1
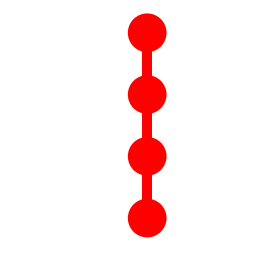
Topological Aspects
- 4 Vertices
- Eigenvalues 0.0000000000
0.5857864618
2.0000000000
3.4142136574
Examples of actual RNAs corresponding to this structure from RNA databases: 152
| PDB_Chain | Type | Discovery Method |
|---|---|---|
| 1A9L_A | PRE-TRNA BULGE-HELIX-BULGE MOTIF | SOLUTION NMR |
| 1CQ5_A | SRP RNA DOMAIN IV | SOLUTION NMR |
| 1CQL_A | SRP DOMAIN IV RNA | SOLUTION NMR |
| 1DRZ_B | RNA (HEPATITIS DELTA VIRUS GENOMIC RIBOZYME) | X-RAY DIFFRACTION |
| 1DUL_B | 4.5 S RNA DOMAIN IV | X-RAY DIFFRACTION |
| 1EHT_A | THEOPHYLLINE-BINDING RNA | SOLUTION NMR |
| 1EXY_A | RNA APTAMER, 33-MER | SOLUTION NMR |
| 1F1T_A | MALACHITE GREEN APTAMER RNA | X-RAY DIFFRACTION |
| 1HQ1_B | 4.5S RNA DOMAIN IV | X-RAY DIFFRACTION |
| 1M5K_B | RNA HAIRPIN RIBOZYME | X-RAY DIFFRACTION |
| 1M5L_A | modified HIV-1 packaging signal stem-loop 1 RNA | SOLUTION NMR |
| 1M5O_B | RNA HAIRPIN RIBOZYME | X-RAY DIFFRACTION |
| 1M5P_B | RNA HAIRPIN RIBOZYME | X-RAY DIFFRACTION |
| 1M5V_B | RNA HAIRPIN RIBOZYME | X-RAY DIFFRACTION |
| 1MZP_B | fragment of 23S rRNA | X-RAY DIFFRACTION |
| 1NBK_A | RNA aptamer | SOLUTION NMR |
| 1NTA_AB | 5'-R(*GP*GP*AP*UP*CP*GP*CP*AP*UP*UP*UP*GP*GP*AP*CP | X-RAY DIFFRACTION |
| 1NTB_AB | 5'-R(*GP*GP*AP*UP*CP*GP*CP*AP*UP*UP*UP*GP*GP*AP*CP | X-RAY DIFFRACTION |
| 1O15_A | THEOPHYLLINE-BINDING RNA | SOLUTION NMR |
| 1P5M_A | 55-MER | SOLUTION NMR |
| 1Q8N_A | RNA Aptamer | SOLUTION NMR |
| 1QZW_F | 7S RNA | X-RAY DIFFRACTION |
| 1SJ3_R | PRECURSOR FORM OF THE HEPATITIS DELTA VIRUS RIBOZY | X-RAY DIFFRACTION |
| 1SJ4_R | PRECURSOR FORM OF THE HEPATITIS DELTA VIRUS RIBOZY | X-RAY DIFFRACTION |
| 1SJF_B | HEPATITIS DELTA VIRUS RIBOZYME | X-RAY DIFFRACTION |
| 1TXS_A | Enteroviral 5'-UTR | SOLUTION NMR |
| 1U3K_A | ABISS7 | SOLUTION NMR |
| 1VBX_B | HEPATITIS DELTA VIRUS RIBOZYME | X-RAY DIFFRACTION |
| 1VBY_B | Hepatitis Delta virus ribozyme | X-RAY DIFFRACTION |
| 1VBZ_B | HEPATITIS DELTA VIRUS RIBOZYME | X-RAY DIFFRACTION |
| 1VC0_B | HEPATITIS DELTA VIRUS RIBOZYME | X-RAY DIFFRACTION |
| 1VC5_B | HEPATITIS DELTA VIRUS RIBOZYME | X-RAY DIFFRACTION |
| 1VC6_B | HEPATITIS DELTA VIRUS RIBOZYME | X-RAY DIFFRACTION |
| 1X9K_DCAB | 5'-R(*UP*CP*GP*UP*GP*GP*UP*AP*CP*AP*UP*UP*AP*CP*CP | X-RAY DIFFRACTION |
| 1YRJ_AB | Bacterial 16 S Ribosomal RNA A Site Oligonucleotid | X-RAY DIFFRACTION |
| 2A9L_A | ABISS7 RNA | SOLUTION NMR |
| 2F4T_BA | 5'-R(P*GP*CP*GP*UP*CP*AP*CP*AP*CP*CP*GP*GP*UP*GP*A | X-RAY DIFFRACTION |
| 2FEY_A | stem-loop IV of Tetrahymena telomerase RNA | SOLUTION NMR |
| 2G5K_AB | 5'-R(*UP*UP*GP*CP*GP*UP*CP*GP*CP*UP*CP*CP*GP*GP*AP | X-RAY DIFFRACTION |
| 2GJW_IJL | 5'-R(*GP*CP*GP*AP*CP*CP*GP*AP*CP*CP*AP*(DU)P*AP*GP | X-RAY DIFFRACTION |
| 2HUA_A | CSFV IRES Domain IIa | SOLUTION NMR |
| 2KTZ_A | HCV IRES Domain IIa RNA | SOLUTION NMR |
| 2KU0_A | HCV IRES Domain IIa RNA | SOLUTION NMR |
| 2KX8_A | 7SK | SOLUTION NMR |
| 2KZL_A | RNA (55-MER) | SOLUTION NMR |
| 2M57_A | RNA_(35-MER) | SOLUTION NMR |
| 2M58_A | RNA (59-MER) | SOLUTION NMR |
| 2MQT_A | RNA (68-MER) | SOLUTION NMR |
| 2MQV_B | RNA (68-MER) | SOLUTION NMR |
| 2N4L_A | RNA (53-MER) | SOLUTION NMR |
| 2N6T_A | RNA (42-MER) | SOLUTION NMR |
| 2N6W_A | RNA (68-MER) | SOLUTION NMR |
| 2NBY_A | IRES RNA (39-MER) | SOLUTION NMR |
| 2NOK_CD | 5'-R(*CP*GP*GP*AP*GP*GP*AP*AP*CP*UP*AP*CP*UP*GP*UP | X-RAY DIFFRACTION |
| 2NOQ_B | 18S ribosomalRNA | ELECTRON MICROSCOPY |
| 2NOQ_DC | 18S ribosomal RNA | ELECTRON MICROSCOPY |
| 2NOQ_E | 25S RIBOSOMAL RNA | ELECTRON MICROSCOPY |
| 2OIH_B | HDV RIBOZYME | X-RAY DIFFRACTION |
| 2OM7_H | Fragment of23S rRNA (H89) | ELECTRON MICROSCOPY |
| 2OM7_J | Fragment of23S rRNA (H76) | ELECTRON MICROSCOPY |
| 2PN3_AB | 5'-R(*CP*GP*GP*AP*GP*GP*AP*AP*CP*UP*AP*CP*UP*GP*UP | X-RAY DIFFRACTION |
| 2PN4_AB | 5'-R(*CP*GP*GP*AP*GP*GP*AP*AP*CP*UP*AP*CP*UP*GP*UP | X-RAY DIFFRACTION |
| 2PXB_B | 4.5 S RNA | X-RAY DIFFRACTION |
| 2PXD_B | 4.5 S RNA | X-RAY DIFFRACTION |
| 2PXE_B | 4.5 S RNA | X-RAY DIFFRACTION |
| 2PXF_B | 4.5 S RNA | X-RAY DIFFRACTION |
| 2PXK_B | 4.5 S RNA | X-RAY DIFFRACTION |
| 2PXL_B | 4.5 S RNA | X-RAY DIFFRACTION |
| 2PXP_B | 4.5 S RNA | X-RAY DIFFRACTION |
| 2PXQ_B | 4.5 S RNA | X-RAY DIFFRACTION |
| 2PXT_B | 4.5 S RNA | X-RAY DIFFRACTION |
| 2PXU_B | 4.5 S RNA | X-RAY DIFFRACTION |
| 2PXV_B | 4.5 S RNA | X-RAY DIFFRACTION |
| 2WW9_D | 25S RRNA | ELECTRON MICROSCOPY |
| 2WWA_D | 25S RRNA | ELECTRON MICROSCOPY |
| 2XKV_B | 4.5S RNA | ELECTRON MICROSCOPY |
| 357D_CBA | RNA (5'-R(*CP*CP*CP*CP*AP*UP*GP*CP*GP*AP*GP*AP*GP* | X-RAY DIFFRACTION |
| 364D_CBA | RNA (5'-R(*CP*CP*CP*CP*AP*UP*GP*CP*GP*AP*GP*AP*GP* | X-RAY DIFFRACTION |
| 3BNL_AB | A site of bacterial ribosome | X-RAY DIFFRACTION |
| 3BNQ_AB | A site of human mitochondrial ribosome | X-RAY DIFFRACTION |
| 3BNR_AB | A site of human mitochondrial ribosome | X-RAY DIFFRACTION |
| 3BNS_AB | A site of human mitochondrial ribosome | X-RAY DIFFRACTION |
| 3J0L_1 | 60S RIBOSOMAL RNA FRAGMENT | ELECTRON MICROSCOPY |
| 3J0L_a | 40S RIBOSOMALRNA FRAGMENT | ELECTRON MICROSCOPY |
| 3J0O_a | 40S ribosomalRNA fragment | ELECTRON MICROSCOPY |
| 3J0P_a | 40S RIBOSOMALRNA FRAGMENT | ELECTRON MICROSCOPY |
| 3J0Q_a | 40S RIBOSOMALRNA FRAGMENT | ELECTRON MICROSCOPY |
| 3JCM_E | SNR14 SNRNA | ELECTRON MICROSCOPY |
| 3JCT_6 | ITS2-1 miscRNA | ELECTRON MICROSCOPY |
| 3LOA_BA | 5'-R(*CP*CP*GP*CP*GP*CP*CP*CP*GP*(5BU)P*CP*AP*CP*A | X-RAY DIFFRACTION |
| 3LQX_B | SRP RNA | X-RAY DIFFRACTION |
| 3OL6_JNKOLP | RNA (5'-R(*AP*AP*GP*UP*CP*UP*CP*CP*AP*GP*GP*UP*CP* | X-RAY DIFFRACTION |
| 3OL9_JNKOLP | RNA (5'-R(*AP*AP*GP*UP*CP*UP*CP*CP*AP*GP*GP*UP*CP* | X-RAY DIFFRACTION |
| 3OLB_JNKOLP | RNA (5'-R(*AP*AP*GP*UP*CP*UP*CP*CP*AP*GP*GP*UP*CP* | X-RAY DIFFRACTION |
| 3SIV_CF | U4atac snRNA | X-RAY DIFFRACTION |
| 3TD0_BA | RNA (5'-R(*UP*UP*GP*CP*GP*UP*CP*GP*CP*GP*(5BU)P*CP | X-RAY DIFFRACTION |
| 3TD1_BA | RNA (5'-R(*UP*UP*GP*CP*GP*UP*CP*GP*CP*GP*(5BU)P*CP | X-RAY DIFFRACTION |
| 3TZR_AB | 5'-R(*CP*GP*AP*GP*GP*AP*AP*CP*UP*AP*CP*UP*GP*UP*CP | X-RAY DIFFRACTION |
| 4ERD_CD | 5'-R(P*GP*GP*UP*CP*GP*AP*CP*AP*UP*CP*UP*UP*CP*GP*G | X-RAY DIFFRACTION |
| 4GPW_AB | RNA (5'-R(*UP*UP*GP*CP*GP*UP*CP*AP*CP*GP*CP*CP*GP* | X-RAY DIFFRACTION |
| 4GPY_AB | RNA (5'-R(*UP*UP*GP*CP*GP*UP*CP*AP*CP*GP*CP*CP*GP* | X-RAY DIFFRACTION |
| 4JRC_B | Distal Stem I region of the glyQS T box leader RNA | X-RAY DIFFRACTION |
| 4K31_BC | RNA (5'-R(*UP*UP*GP*CP*GP*UP*CP*GP*UP*UP*CP*CP*GP* | X-RAY DIFFRACTION |
| 4OJI_A | RNA (52-MER) | X-RAY DIFFRACTION |
| 4P3S_BA | 5'-D(*UP*UP*GP*CP*GP*UP*CP*CP*CP*GP*(5BU)P*CP*GP*A | X-RAY DIFFRACTION |
| 4P3T_AB | 5'-D(*UP*UP*GP*CP*GP*UP*CP*CP*CP*GP*(5BU)P*CP*GP*A | X-RAY DIFFRACTION |
| 4P3U_BA | bacterial A1408U-mutant ribosomal decoding site | X-RAY DIFFRACTION |
| 4P43_BA | bacterial A1408U-mutant ribosomal decoding site | X-RAY DIFFRACTION |
| 4PDB_I | SELEX RNA aptamer | X-RAY DIFFRACTION |
| 4PDQ_AB | RNA (5'-*UP*UP*GP*CP*GP*UP*CP*AP*CP*GP*CP*CP*GP*GP | X-RAY DIFFRACTION |
| 4PHY_AB | RNA (26-MER) | X-RAY DIFFRACTION |
| 4PKD_V | U1 snRNA stem-loops 1 and 2 (55-MER) | X-RAY DIFFRACTION |
| 4PMI_A | Rev-Response-Element RNA | X-RAY DIFFRACTION |
| 4PR6_B | HDV RIBOZYME SELF-CLEAVED | X-RAY DIFFRACTION |
| 4PRF_B | HEPATITIS DELTA VIRUS RIBOZYME | X-RAY DIFFRACTION |
| 4RGE_A | ENV22 TWISTER RIBOZYME | X-RAY DIFFRACTION |
| 4RGF_B | ENV22 TWISTER RIBOZYME | X-RAY DIFFRACTION |
| 4TS0_YX | Spinach RNA aptamer, bimolecular construct | X-RAY DIFFRACTION |
| 4V8Y_CN | EUKARYOTIC RIBOSOMAL L1_RRNA | ELECTRON MICROSCOPY |
| 4WFA_Y | 5S RRNA | X-RAY DIFFRACTION |
| 4WFB_Y | 5S RRNA | X-RAY DIFFRACTION |
| 4ZNP_A | pfI Riboswitch | X-RAY DIFFRACTION |
| 5AXW_B | RNA (73-MER) | X-RAY DIFFRACTION |
| 5BTM_A | RNA (55-mer) | X-RAY DIFFRACTION |
| 5BWS_AB | RNA (5'-D(*UP*UP*GP*CP*GP*UP*CP*AP*CP*GP*CP*CP*GP* | X-RAY DIFFRACTION |
| 5BXK_CD | RNA (5'-D(P*UP*UP*GP*CP*GP*UP*CP*AP*CP*GP*CP*CP*GP | X-RAY DIFFRACTION |
| 5CZZ_B | RNA (73-MER) | X-RAY DIFFRACTION |
| 5EW4_BA | RNA (5'-R(*CP*CP*CP*CP*GP*GP*CP*CP*CP*CP*GP*GP*CP* | X-RAY DIFFRACTION |
| 5EW7_BA | RNA (5'-R(*CP*CP*CP*CP*GP*GP*CP*CP*CP*CP*GP*GP*CP* | X-RAY DIFFRACTION |
| 5GAD_1 | ESRP 4.5S RNA | ELECTRON MICROSCOPY |
| 5GAF_1 | SRP 4.5S RNA | ELECTRON MICROSCOPY |
| 5GAG_1 | SRP 4.5S RNA | ELECTRON MICROSCOPY |
| 5GAH_1 | SRP 4.5S RNA | ELECTRON MICROSCOPY |
| 5H1S_C | SPINACH CHLOROPLAST 4.5S RRNA | ELECTRON MICROSCOPY |
| 5ND9_B | 5S ribosomal RNA | ELECTRON MICROSCOPY |
| 5OB3_A | RNA aptamer (69-MER) | X-RAY DIFFRACTION |
| 5T7V_C | 5S ribosomal RNA | ELECTRON MICROSCOPY |
| 5TF6_B | U6 snRNA | X-RAY DIFFRACTION |
| 5U30_B | SGRNA | X-RAY DIFFRACTION |
| 5U31_B | sgRNA | X-RAY DIFFRACTION |
| 5U34_B | SGRNA | X-RAY DIFFRACTION |
| 5UNE_AB | RNA (47-MER) | X-RAY DIFFRACTION |
| 5X2G_B | sgRNA | X-RAY DIFFRACTION |
| 5X2H_B | SGRNA | X-RAY DIFFRACTION |
| 5XZ1_AB | RNA (5'-R(*UP*GP*CP*GP*UP*CP*GP*CP*UP*CP*CP*GP*GP* | X-RAY DIFFRACTION |
| 5Z1H_BA | RNA (5'-R(*UP*UP*GP*CP*GP*UP*CP*AP*CP*GP*CP*CP*GP* | X-RAY DIFFRACTION |
| 5ZEG_ABC | RNA (5'-R(*UP*UP*GP*CP*GP*UP*CP*(1MA)P*CP*GP*UP*CP | X-RAY DIFFRACTION |
| 5ZEJ_AB | RNA (5'-R(P*UP*GP*CP*GP*UP*CP*(1MA)P*CP*GP*UP*CP*G | X-RAY DIFFRACTION |
| 6CF2_G | RNA (35-MER) | X-RAY DIFFRACTION |
| 6D90_4 | CrPV-IRES | ELECTRON MICROSCOPY |
| 6ELZ_6 | Internal transcribed spacer 2 | ELECTRON MICROSCOPY |
| 6EM1_6 | 5.8S ribosomal RNA | ELECTRON MICROSCOPY |